Ants, Bees, Genomes & Evolution @ Queen Mary University London
Published: 25 April 2019
Choosing the Best Gene Predictions with GeneValidator
Ismail Moghul, Anurag Priyam, Yannick Wurm
Gene Prediction. Methods in Molecular Biology, Volume 1962, Chapter 16
Abstract
GeneValidator is a tool for determining whether the characteristics of newly predicted protein-coding genes are consistent with those of similar sequences in public databases. For this, it runs up to seven comparisons per gene. Results are shown in an HTML report containing summary statistics and graphical visualizations that aim to be useful for curators. Results are also presented in CSV and JSON formats for automated follow-up analysis.
Here, we describe common usage scenarios of GeneValidator that use the JSON output results together with standard UNIX tools. We demonstrate how GeneValidator’s textual output can be used to filter and subset large gene sets effectively. First, we explain how low-scoring gene models can be identified and extracted for manual curation—for example, as input for genome browsers or gene annotation tools. Second, we show how GeneValidator’s HTML report can be regenerated from a filtered subset of GeneValidator’s JSON output. Subsequently, we demonstrate how GeneValidator’s GUI can be used to complement manual curation efforts. Additionally, we explain how GeneValidator can be used to merge information from multiple annotations by automatically selecting the higher-scoring gene model at each common gene locus. Finally, we show how GeneValidator analyses can be optimized when using large BLAST databases.
1. Introduction
Using accurate gene annotations is important because they affect subsequent analyses [1]. For some species, annotations can be downloaded directly from a public database such as Ensembl or NCBI [2]. For newly sequenced species, approaches to identify protein-coding genes in a genome sequence typically combine evidence from multiple data sources (including ab initio models, ESTs, RNA-seq, and protein alignments) [3–5]. Whether gene feature annotations are downloaded from a public database or are newly generated, they may contain errors resulting from biases of the underlying data, algorithmic choices [6], and the general limitations of a one-dimensional representation of DNA sequences. Common errors include frameshifts, incorrect exon–intron structure, incorrect merging of adjacent genes, and incorrect splitting of genes at long intron positions [7].
Fig.1 High-level schematic of the steps carried out by GeneValidator
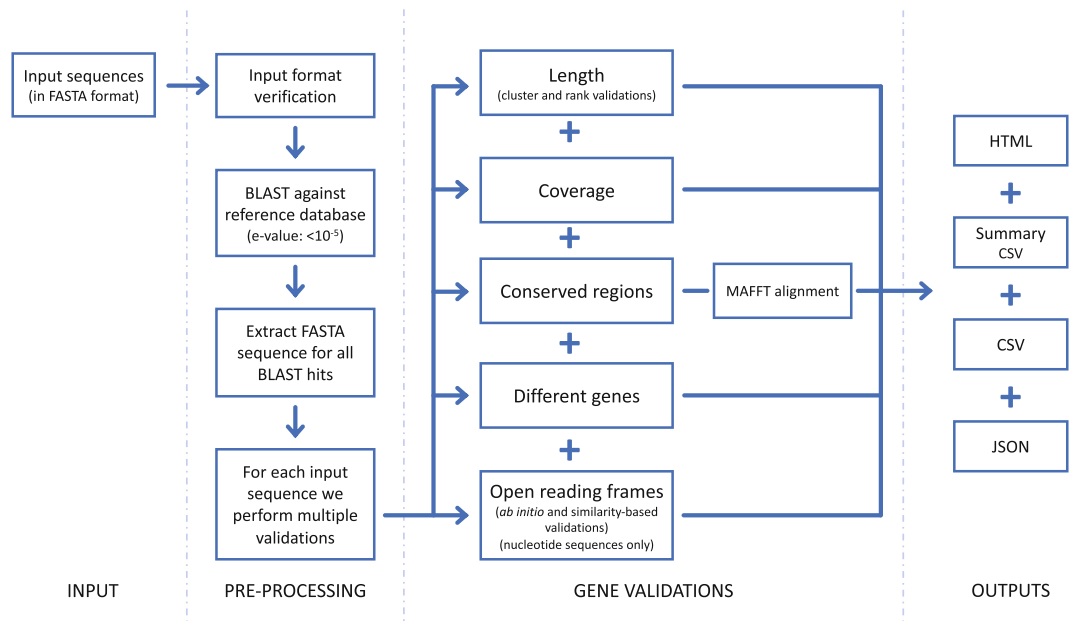
We previously described GeneValidator (GV), a tool to evaluate the quality of protein-coding gene predictions based on comparisons with a database of known proteins [8]. In brief (Fig. 1), GV first runs a BLAST search against the given database, retaining sequences of hits with e-value stronger than 10-5. Next, GV runs up to seven validations on each gene prediction. Each validation tests if the characteristics of the query gene deviate from those of similar sequences in the reference database. Based on predefined thresholds, the result of each validation is a pass or a fail. The overall score of the prediction is a scaled percentage of the validations that passed. Predictions with a score lower than 75 (i.e., more than one failed validation) may be regarded as potentially problematic. Explanation of the approach and an overview of the data underlying each validation are included in the HTML report, along with several visualizations to facilitate interpretation. Detailed results are also available in CSV and JSON format for spreadsheet and programmatic access.
Results produced by GV depend on the quality and coverage of the database used for validation. Furthermore, higher scores indicate consistency with database sequences and not biological truths. Several publicly available databases of protein sequences such as Swiss-Prot [9], UniRef50 [9, 10], TrEMBL [9], or NR [2] can be used with GV. The GV approach becomes increasingly reliable as proteomes of more species are submitted to these databases by the global research community, and as the qualities of submitted sequences improve due to experimental validation, manual verification by experts, and technological and algorithmic advances in sequencing and automated gene prediction. We created GV to be flexible. Many of GV’s features are designed to facilitate automatic processing of large gene sets (e.g., whole-genome annotation) as part of custom workflows. These include GV’s versatile JSON output, ability to leverage HPC facilities, and the possibility to use advanced BLAST search options. GV also includes a web server that can be used as a shared resource. Here, we discuss five common use cases of GV that can be easily incorporated into custom workflows.
2. Installing and Running GeneValidator
GV runs on Linux and macOS. To install GV, run the command shown below. This will install GVand all its dependencies to a directory called “genevalidator” in the current working directory.
sh -c "$(curl -fsSL
https://install-genevalidator.wurmlab.com)"
The software includes example sequences to test the installation. The following command can be used to run GV on these example sequences with the included Swiss-Prot database. GV will print the results of validations for each gene prediction to the terminal, ending with a summary, and the directory where detailed results were saved to.
genevalidator --db genevalidator/blast_db/swissprot \
--num_threads 4 \
genevalidator/exemplar_data/protein_data.fa
3. GeneValidator Workflows
A gene set will almost inevitably contain some gene predictions with low scores. It can be desirable to curate these manually. Here, we begin by providing two approaches to facilitate inspection of these low-scoring predictions. First (Subheading 3.1), we show how to use GV’s JSON output to extract the sequence identifiers of low-scoring gene predictions. Among other things, these can be used to subset the initial gene set, to prioritize inspection in a genome browser [11], or for annotation editing in a tool such as Apollo [12]. Second (Subheading 3.2), we show how to create a new HTML report by subsetting GV’s JSON output. This can reduce the need to navigate through a long HTML report. Subsequently (Subheading 3.3), we introduce GV’s graphical interface. This is helpful for rapidly viewing how GV’s validation results change during manual curation.
We also provide guidance on two more general challenges based on our applications of GV. First (Subheading 3.4), we show how GV can be used to automatically select the best gene model from multiple gene sets at each common gene locus. Furthermore (Subheading 3.5), we show how to restrict GV to use a specific subset of a BLAST database. This is to avoid BLAST searching against sequences unlikely to be informative.
3.1. Extracting Sequence Identifiers of Low-Scoring Gene Predictions
GV’s JSON output can be used with JQ (https://stedolan.github.io/jq/), a command-line JSON processor (included in the GV package), to select gene predictions matching a particular criterion and access validation results and associated metadata. In the example below, we extract identifiers of predictions with a score lower than 75 (i.e., having failed more than one validation) and having at least two BLAST hits for manual curation. The idea is that while having two BLAST hits is insufficient for GV’s statistical tests (and thus results in a low score), they may provide sufficient evidence for biologically interpreting whether the prediction could be appropriate.
- Extract FASTA header of gene predictions that have more than two BLAST hits and an overall score of less than 75.
jq --raw-output ".[] | select(.no_hits >= 2 and .overall_score < 75) | .definition" input_file_results.json \ > sequence_definitions.txt - Extract sequence identifier (first word of the FASTA header) using the cut command.
cut -d " " -f 1 sequence_definitions.txt \ > sequence_ids.txt
3.2. Subsetting the HTML Report to Only Low-Scoring Gene Predictions
GV’s JSON output can be filtered using JQ and input back to GV to reproduce results for the selected gene predictions. This is useful to create smaller HTML reports, for example, focusing on a particular gene family. In the example below, we subset GV’s output for the low-scoring gene predictions selected in Subheading 3.1.
- Select gene predictions that have more than two BLAST hits and an overall score of less than 75.
jq "[ .[] | select(.no_hits >= 2 and .overall_score < 75) ] | sort_by(.overall_score)" input_file_results.json \ > input_file_results_subset.json - Reproduce GV’s output.
genevalidator --json input_file_results_subset.json
3.3. Using GeneValidator Web Server to Iteratively Refine Gene Models
Although running GV from the command line is ideal for processing of large datasets and custom workflows, a graphical user interface can facilitate iterative usage. For example, during manual curation of gene models, running GV repeatedly as a gene model is revised can help a curator verify that changes they are making indeed improve the gene model. Building on the lessons learnt when developing the SequenceServer BLAST interface [13], we also built a graphical user interface (app) for GV that is accessible through a web browser.
- Launching GV app requires the path to a directory containing one or more BLAST databases; the interface (accessible at
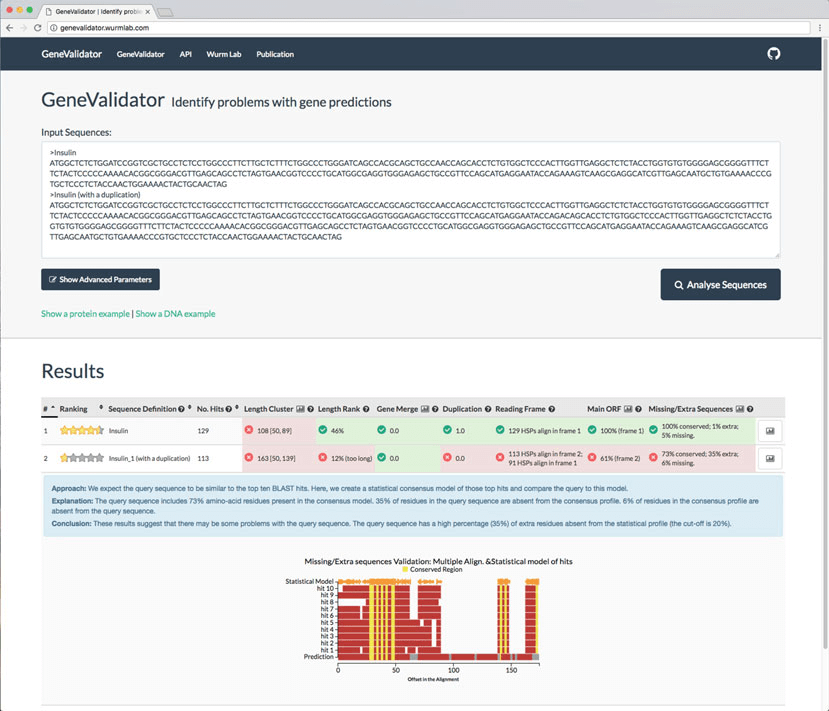
http://localhost:5678) is opened automatically in the default browser.genevalidator app --num_threads 4 \ --database_dir genevalidator/blast_db/ - To validate gene predictions, paste the corresponding FASTA sequences into the text area, select the database to compare to, and click “Analyse Sequences” (Fig. 2). The results are then shown on the same page. We also host a GV web server at https://genevalidator.wurmlab.com with two caveats: first, it is suitable for up to ten queries at a time, and second, given computational constraints on this server, we only provide the Swiss-Prot and the UniRef50 databases.
3.4. Merging Gene Predictions from Two Different Sources
Different gene prediction approaches are unlikely to generate identical gene models for a locus. GV can be used to select the higherscoring gene model for each locus from multiple gene sets. Briefly, we first identify annotations corresponding to the same locus from the different sources (steps 1–3 below). Subsequently, we generate a FASTA file containing alternative predictions for each locus and use GV’s --select_single_best option to select the higher scoring one (step 4 below).
We make multiple simplifying assumptions to generate a mapping of annotations corresponding to the same locus from the different sources (steps 1–3 below). Specifically, we assume that we have a single transcript (splice form) per source per locus, that gene predictions from different loci do not overlap, and that annotations are available in a GFF3 format file. Often, additional preprocessing of gene sets will be necessary to fulfill these assumptions.
Fig.2 A screenshot of the GeneValidator web application as launched from the command line via genevalidator app or by accessing genevalidator.wurmlab.com
- Intersect the transcript annotations in the GFF3 files (requires prior installation of bedtools). We require that both hits are on the same strand (
-s). If comparing more than two GFF3 files, see the bedtools documentation (-bcan take multiple values). The output file contains the entire input record from both input files (-wa -wb).awk ’/\tmRNA\t/’ geneset1.gff > geneset1_mrnas.gff awk ’/\tmRNA\t/’ geneset2.gff > geneset2_mrnas.gff bedtools intersect -wa -wb -s \ -a geneset1_mrnas.gff -b geneset2_mrnas.gff \ > geneset_overlaps.bed - Extract the GFF3 attributes columns (i.e., the 9th and 18th column) which contain the sequence identifiers.
awk ’{printf ("%s;\t%s;\n", $9, $18)}’ \ geneset_overlaps.bed > attributes_columns.tsv - Extract the sequence identifiers from the attributes columns.
perl -nle ’@ids = /ID=(.*?);/g; print join("\t", @ids) if @ids’ \ attributes_columns.tsv > mapping_ids.tsv - Now that we have identifiers of the annotations corresponding to the same locus from both the gene sets, their respective sequences can be extracted and then used with GV’s
--select_single_bestoption.
(a) Create indexes for each of the FASTA files (requires prior installation of samtools).
samtools faidx geneset1.fasta
samtools faidx geneset2.fasta
(b) Create output FASTA file.
touch output.fa
(c) Loop over the mapping_ids.tsv file. Extract FASTA sequence for each ID, and write them to a temporary FASTA file. Run GV using the --select_single_best option on the temporary FASTA file. The --select_single_best mode prints the highest-scoring sequence to STDOUT in FASTA format, which is written to the output file previously created.
cat mapping_ids.tsv | while read -r line; do
echo "$line" | cut -f 1 | \
xargs samtools faidx geneset1.fasta \
> gv_run_tmp.fa
echo "$line" | cut -f 2 | \
xargs samtools faidx geneset2.fasta \
>> gv_run_tmp.fa
genevalidator --select_single_best gv_run_tmp.fa \
>> output.fa
rm gv_run_tmp.fa
done
It may be desirable to include gene models unique to both sets in the final output. We leave this as an exercise for the reader.
3.5. Using NCBI’s Nonredundant Database of Protein Sequences with GV
While it is desirable to validate gene predictions against a gold standard database like Swiss-Prot, its limited coverage [9] makes this challenging for many species. At the same time, technological advances continue to increase the quality of automated predictions [14]. This makes it tempting to use a more comprehensive database such as NCBI’s nonredundant collection (NR) of manually reviewed as well as automatically generated protein sequences for validation. However, the large size of the NR database means BLAST searches can take days. We show how to use BLAST’s ability to restrict searches to a list of identifiers [15] to accelerate a GV analysis. For this, we first restrict the BLAST search to a particular taxonomic lineage to avoid BLAST searching against sequences unlikely to be informative. Additionally, we exclude sequences from the focal species to avoid circular self-validation.
For the implementation below, we consider the example of the red fire ant, Solenopsis invicta [16]. We first obtain taxon identifiers of all species in Eukaryota (id: 2759). Subsequently, we exclude all Solenopsis species (taxonomy id: 13685). We then obtain GenInfo identifiers (GI numbers) of all sequences in the retained taxa. We finally run GV using this list.
- Obtain a list of eukaryotic taxon identifiers (this requires prior installation of Taxonkit [17]).
taxonkit list --ids 2759 --indent "" \ > taxon_ids_eukaryotes.txt - Obtain a list of Solenopsis taxon identifiers.
taxonkit list --ids 13685 --indent "" \ > taxon_ids_solenopsis.txt - Subtract the two.
grep -Fvx -f taxon_ids_solenopsis.txt \ taxon_ids_eukaryotes.txt > taxon_ids.txt - Download a tab-delimited file from NCBI linking taxon ids and GI Numbers.
curl -L -O ftp://ftp.ncbi.nih.gov/pub/taxonomy/accession2taxid/prot. accession2taxid.gz - Use csvtk (https://github.com/shenwei356/csvtk), a multithreaded CSV/TSV processor (packaged with GV), to extract the rows where the taxid is in the taxon_ids.txt file.
zcat prot.accession2taxid.gz | \ csvtk --tabs grep --fields taxid \ --pattern-file taxon_ids.txt | \ cut -f 4 | tail -n +2 > gi_list.txt - Finally, we pass this file to GV using
--blast_optionoption.genevalidator --blast_options "-gilist gi_list.txt" \ --db nr --num_threads 40 geneset1.fa
Starting with BLASTþ version 2.8.0 (in development at the time of this writing), steps 4 and 5 can be skipped, and the list of taxon ids from step 3 can be passed directly to BLAST using the new -taxidlist option.
4. Tips and Tricks
-
GV’s overall score is based on the percentage of validations that pass, i.e., where the score is above a threshold that we have determined to be appropriate. To emphasize the fact that GV results are highly dependent on the quality of information in databases and cannot be solely relied upon to classify a “perfect” gene prediction, the overall score is decreased by 10%. The highest possible score is thus 90%.
-
GV will run the validations provided there are at least five BLAST hits for a given prediction. This can be changed using the
--min_blast_hitsoption. A higher number of BLAST hits will increase the relevance of the comparisons. -
GV generates several summary statistics for the input gene set. These include first, second, and third quartiles of the overall scores, number of good and bad predictions, and number of predictions with insufficient BLAST hits. In addition to providing an overview of the quality of the input gene set, the summary statistics can be used to choose between predictions from two different sources.
-
GV includes a tool for downloading sequence databases from NCBI to use for comparisons (i.e.,
genevalidator ncbiblast-dbs). This is a parallelized alternative to the “update_blastdb.pl” script included in BLASTþ package. -
GV is also able to run BLAST searches on NCBI servers using BLAST’s
-remoteoption (e.g.,enevalidator --db `swissprot -remote` geneset.fa). This has the benefit of being able to immediately use the most up-to-date version of a given database. However, using a remote BLAST database is very slow. We recommended using this for validating only a few genes (e.g., fewer than 25). -
It is possible to run BLAST independently and to subsequently provide the output XML (
-outfmt 5) or tab-delimited (- outfmt 6) to GV. This can be particularly useful if BLAST results have already been produced for other analyses or when BLAST can be run on a cluster. -
BLAST is often the slowest step of GV pipeline, especially when working with large datasets. In such cases, DIAMOND [18] can be used instead of BLAST for (up to 20,000!) faster database searching. Since DIAMOND’s XML output is compatible with BLAST, it can be used directly with GV along with one additional input, i.e., a FASTA file of hit sequences (when used with BLAST, GV is able to automatically extract hit sequences from BLAST database). Our wiki (https://github.com/wurmlab/genevalidator/wiki) provides detailed instructions for using GV with DIAMOND.
-
To resume a terminated analysis, GV can be run with
-- resumeoption. In resume mode, GV skips previously successful steps, including running BLAST. Gene predictions that were successfully processed are skipped as well. -
It is possible to split an input gene set into multiple chunks, run GV on each chunk across multiple compute nodes, and combine the results for each chunk into a single report. (a) After splitting the input file and running GV on each input file, the following command can be used to merge the individually produced GV JSON files.
cat */*.json | jq ".[]" | jq --slurp "." > MERGED_JSON
(b) The merged JSON can then be used to produce a single report for the whole gene set.
genevalidator --json MERGED_JSON
References
-
Yandell M, Ence D (2012) A beginner’s guide to eukaryotic genome annotation. Nat Rev Genet 13:329–342
-
Benson DA, Cavanaugh M, Clark K, KarschMizrachi I, Ostell J, Pruitt KD et al (2018) GenBank. Nucleic Acids Res 46:D41–D47
-
Holt C, Yandell M (2011) MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12:491
-
Hoff KJ, Lange S, Lomsadze A, Borodovsky M, Stanke M (2016) BRAKER1: unsupervised RNA-Seq-based genome annotation with GeneMark-ET and AUGUSTUS. Bioinformatics 32:767–769
-
Keilwagen J, Hartung F, Paulini M, Twardziok SO, Grau J (2018) Combining RNA-seq data and homology-based gene prediction for plants, animals and fungi. BMC Bioinformatics 19:189
-
Schnoes AM, Brown SD, Dodevski I, Babbitt PC (2009) Annotation error in public databases: misannotation of molecular function in enzyme superfamilies. PLoS Comput Biol 5:e1000605
-
Steijger T, Abril JF, Engstro¨m PG, Kokocinski F, RGASP Consortium, Hubbard TJ et al (2013) Assessment of transcript reconstruction methods for RNA-seq. Nat Methods 10:1177–1184
-
Dra˘gan M-A, Moghul I, Priyam A, Bustos C, Wurm Y (2016) GeneValidator: identify problems with protein-coding gene predictions. Bioinformatics 32(10):1559–1561
-
The UniProt Consortium (2017) UniProt: the universal protein knowledgebase. Nucleic Acids Res 45:D158–D169
-
Suzek BE, Wang Y, Huang H, McGarvey PB, Wu CH, The UniProt Consortium (2015) UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31:926–932
-
Buels R, Yao E, Diesh CM, Hayes RD, MunozTorres M, Helt G et al (2016) JBrowse: a dynamic web platform for genome visualization and analysis. Genome Biol 17:66
-
Lee E, Helt GA, Reese JT, Munoz-Torres MC, Childers CP, Buels RM et al (2013) Web Apollo: a web-based genomic annotation editing platform. Genome Biol 14:R93
-
Priyam A, Woodcroft BJ, Rai V, Munagala A, Moghul I, Ter F et al (2015) Sequenceserver: a modern graphical user interface for custom BLAST databases. bioRxiv. https://doi.org/10.1101/033142
-
Minoche AE, Dohm JC, Schneider J, Holtgr€awe D, Vieho¨ver P, Montfort M et al (2015) Exploiting single-molecule transcript sequencing for eukaryotic gene prediction. Genome Biol 16:549
-
Bethesda (MD): National Center for Biotechnology Information (2008) BLAST® Command Line Applications User Manual [Internet] - Limiting a Search with a List of Identifiers. https://www.ncbi.nlm.nih.gov/books/NBK279673. Accessed 13 Sept 2018
-
Wurm Y, Wang J, Riba-Grognuz O, Corona M, Nygaard S, Hunt BG et al (2011) The genome of the fire ant Solenopsis invicta. Proc Natl Acad Sci U S A 108 (14):5679–5684
-
Shen W, Xiong J (2019) TaxonKit: a crossplatform and efficient NCBI taxonomy toolkit. bioRxiv. https://doi.org/10.1101/513523
-
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60
Acknowledgments
This work was supported by the Natural Environment Research Council [grant NE/L00626X/1] and the Biotechnology and Biological Sciences Research Council [grant BB/K004204/1 and BB/M009513/1]. This research used Queen Mary’s Apocrita HPC facility, supported by QMUL Research-IT https://doi.org/10.5281/zenodo.438045.
Author information
Ismail Moghul and Anurag Priyam contributed equally to this work.
Authors and Affiliations
UCL Cancer Institute, University College London, London, UK
Ismail Moghul
School of Biological and Chemical Sciences, Queen Mary University of London, London, UK
Anurag Priyam & Yannick Wurm
Corresponding authors
Correspondence to Ismail Moghul or Yannick Wurm.